Summary
Based on data collected over the last year from various forensic labs, (see http://www.cstl.nist.gov/biotech/strbase/mixture.htm) the majority of DNA evidence mixtures observed contain material from two or three different sources. Data sets containing two- or three-person mixtures, generated in-house or obtained from collaborators, are being evaluated with different software programs to determine optimal ranges for reliable mixture interpretation.
UPDATE
Results from the 2005 and 2013 DNA Mixtures Interlaboratory Studies have been published.
Description
Mixtures of DNA from more than one source are commonly seen in forensic evidentiary samples and are challenging to interpret. Over the past decade, our group at NIST has conducted several interlaboratory studies involving mixture interpretation with short tandem repeat (STR) markers in order to better characterize issues that arise when trying to decipher the contributors to a DNA mixture. In addition, several software programs have been evaluated for their ability to aid in mixture interpretation.
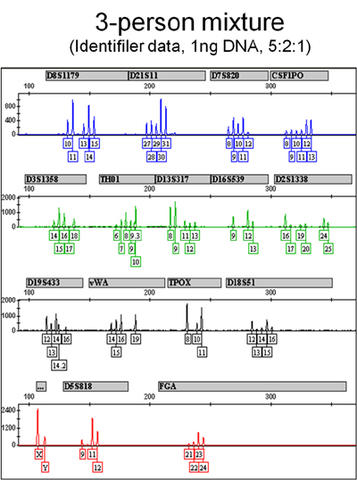
Figure 1. Electropherogram of a 3-person mixture using Applied Biosystem's Identifiler STR kit with 1ng total input DNA at 5:2:1 ratio of the components.
Major Accomplishments
- Collected 15 years of data from interlaboratory studies (see http://www.cstl.nist.gov/biotech/strbase/interlab.htm )
- Examined mixture data sets from collaborators including US Bureau of Alcohol, Tobacco, Firearms, and Explosives (ATF) and National Institute of Justice (NIJ) Expert Systems Testbed (NEST) Project as well as data generated in-house at NIST
- Evaluated software including i-STREAM portion of FSS-i3 v4.1.3 (Promega), the Web-based Least Squares Deconvolution (Web-LSD) and the Excel-based program DNA_DataAnalysis v2.1.3 (USACIL)

